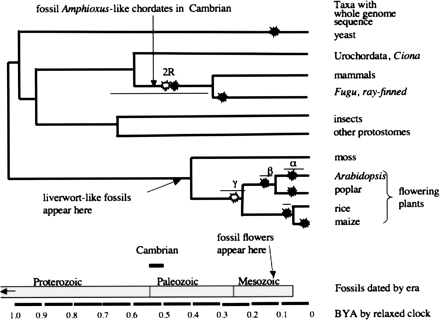
Paleopolyploidy in eukaryotes plotted onto a phylogenetic tree (relaxed clock by Douzery et al. 2004) of eukaryotes using species that have a majority of genome sequenced. Tetraploidy events are denoted with a black starburst, and large-scale segmental events (possible tetraploidies) with an outlined starburst. Outlined starbursts and times previous are in the “twilight zone” (Simillion et al. 2004). Each event has a range of suggested time points indicated by the length of the thin line. The geological eras (Freeman and Herron 2004) are indicted by rectangles. The position of amphioxus is inferred from Hox gene research (Furlong and Holland 2004). The first appearance of a liverwort-like plant reflects a recent fossil find (Wellman et al. 2003). The angiosperm fossil record and first appearance of flowers in the Cretaceous have been reviewed (Friis et al. 2005); note the huge discrepancy between fossil-based and relaxed clock ages for the appearance of flowering plants. Major tetraploidy references include for yeast (Wolfe and Shields 1997; Kellis et al. 2004); chordates (McLysaght et al. 2002; Simillion et al. 2004); ray-finned fish (see Taylor et al. 2003); Arabidopsis α, β γ, identified with Greek letters on the tree (Bowers et al. 2003) using a comparative gene-tree approach; Maere et al. (2005) used a molecular clock to deduce three similar events called 1R, 2R, and 3R, and general Arabidopsis large-scale duplication (Blanc et al. 2000; Ku et al. 2000; Simillion et al. 2002); poplar (D. Rokhsar, Joint Genomes Institute, DOE, >60 Mya; pers. comm.); rice (Paterson et al. 2005; Tian et al. 2005; Yu et al. 2005), and maize (Gaut and Doebley 1997). (BYA) Billion years ago.











