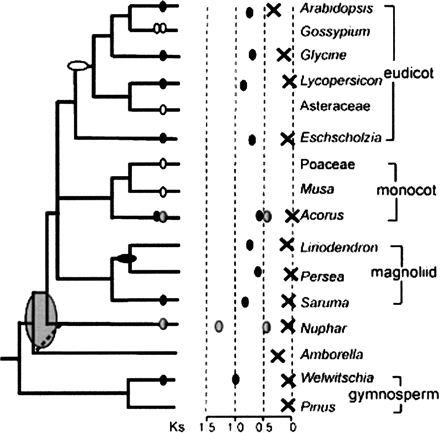
Phylogenetic summary of paleopolyploidy events estimated by the mixture model approach and their distribution among angiosperm and gymnosperm lineages. Scaled graph in center with Xs corresponding to median Ks of pairs from background gene duplications, while small ovals indicate the median Ks of possible concentrated duplications in the history of particular lineages. The phylogenetic tree at left shows the likely placement of detected genome-scale duplications. Uncertainty in phylogenetic timing of what may be a single duplication event at the base of the angiosperms is indicated with a wide oval that covers possible branching points compatible with the Ks evidence. Hollow ovals indicate duplications identified in previous studies using paralogous genes or genomic data from those lineages.











