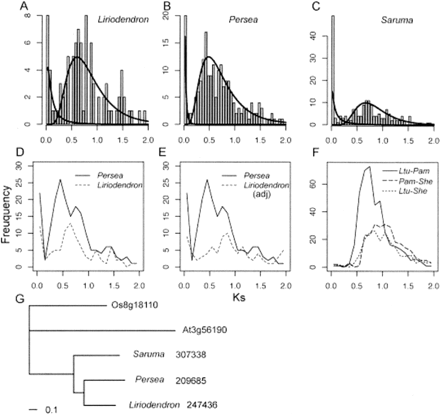
Ks distributions of paralogs and orthologs among magnoliids suggest independent duplications and possibly shared genome duplication events in Laurales (Persea) and Magnoliales (Liriodendron). (A,B,C) The Ks distributions for (A) Liriodendron, (B) Persea, and (C) Saruma, with fitted lines based on the mixture model analysis. (D) The Ks distribution for Liriodendron and Persea, without scaling for rate differences between lineages. (E) Ks distribution for paralogs in Liriodendron after rate calibration (adj = adjusted), compared with that of Persea, suggesting recent independent duplication and older shared genome-scale duplications. (F) Ks distribution for orthologs of two magnoliid species. (Ltu) Liriodendron; (Pam) Persea; (She) Saruma. (G) Phylogeny of one representative orthologous gene set used for relative rate estimates. The branch lengths show the estimated relative rates of synonymous evolution in respective species.











