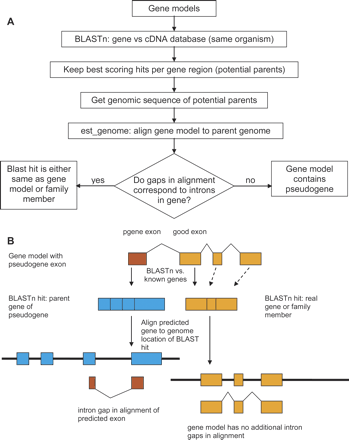
The intron location method of pseudogene finding. (A) Flow diagram of the method. See text for details. (B) All predicted gene models are used for BLASTn against a database of known genes. When a pseudogene is incorporated in the gene model, it will hit its parent gene in the BLAST search (left side of diagram). Alignment with the genomic location of the parent gene will usually show intron gaps. Gene model segments that are not derived from pseudogenes may hit a family member elsewhere in the genome (right side of diagram). In this case, alignment of the prediction to the genomic region of the parent will typically include gaps where introns are predicted in the gene model.











