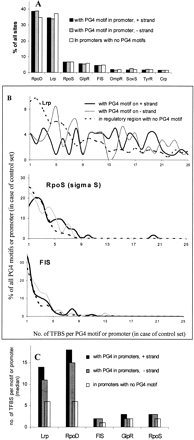
Global regulators Lrp, FIS, and GlpR and sigma factors σ70 and σS are predominantly associated with PG4 motifs in Escherichia coli. We computationally mapped target sites for 55 DNA-binding proteins in the region flanking (100 bp) PG4 motifs present within −200 bp of start codons in the + strand (118 motifs) and − strand (96 motifs). Sites were also mapped to 445 promoter regions (within −200 bp of start codon) devoid of PG4 motifs as a control set. (A) Overall representation of sites (for nine factors with >1% sites) as a percentage of total sites for 55 DNA-binding proteins is shown for the respective regions. (B) Frequency distribution of TFBS. Motifs or promoters (%) were plotted against the number of sites found either flanking the motifs or within the promoter (in case of control set); representative plots for three factors are shown (for others, see Supplemental Fig. S6). Distributions were observed to be significantly (P < 0.001) different for Lrp, RpoD, FIS, RpoS, and GlpR when compared between the + or − strand and the control set, while SoxS, TyrR, Crp, and OmpR did not show a statistically significant difference (P > 0.05). (C) Target sites (median) per motif (+/− strand) or promoter (control set) are shown for five factors with significantly different distribution. Nonparametric comparisons were done using the Mann-Whitney U-test; the P-values for respective comparisons are shown in Supplemental Table S8.











