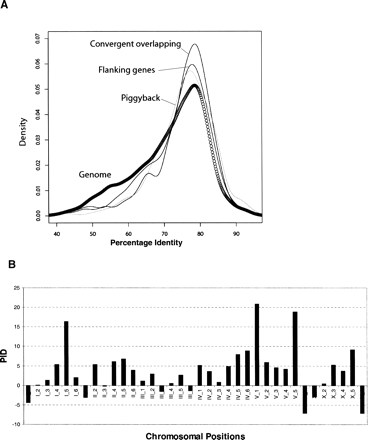
Conservation of overlapping genes. (A) Distribution of protein percentage identity between C. elegans and C. briggsae orthologous for overlapping genes (piggyback, convergent overlapping genes, and flanking genes of the opposite-strand nested gene pairs) and genes in the whole C. elegans genome. (B) Gene conservation in different genomic divisions. Each chromosome (I, II, III, IV, V, X) is divided into six bins (e.g., I_1, I_2, . . . , X_5, X_6). Each bar represents the averaged protein percentage identity for overlapping genes subtracted by that of all other genes in the same bin. Positive bars indicate that overlapping genes are more conserved than the other genes.











