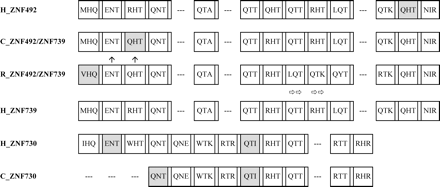
Zinc finger alignment for the loci in the ZNF492 clade. The alignment includes the human paralogs and their orthologs if found in chimpanzee and rhesus genomic data. Finger boxes, amino acid codes, degenerate finger shading, and frameshifts are as in Figure 4B. The chimpanzee ZNF492/ZNF739 homolog finger-exon sequence is in a location and orientation matching human ZNF739 in the panTro1 draft, but there are assembly gaps around it and in the location where ZNF492 (if both are present) would be located. Newman et al. (2005) recorded a possible inversion in this region between human and chimpanzee. The location of the rhesus copy is unknown. Therefore, the presumed nonhuman ortholog sequences are not explicitly identified with either human locus name. The horizontal arrows below the rhesus ZNF492/ZNF739 protein homolog represent a finger in the protein alignment that is divergent from the same-numbered finger in the human and chimpanzee homologs. The rhesus ZNF492/ZNF739 protein’s fingers 8 and 9 are similar enough to fingers 10 and 11 that in this case, as an alternative to mutational change in the finger amino acid sequence, the pattern in this species could also be explained by a loss of fingers followed by an internal duplication of the aforementioned finger motifs, restoring the array to the same number of zinc finger motifs. The ZNF730 gene contains a stop codon after the 12th finger, eliminating several 3′-end ZNF motifs that are included in the ZNF492 and ZNF739 proteins. For chimpanzee ZNF730, the available genomic sequence is missing the first three finger motifs, but the cross-primate PCR results showed a similar band for chimpanzee and human (data not shown).











