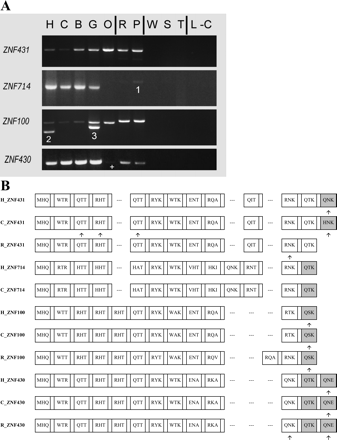
(A) PCR results using primers for selected members of the ZNF431 clade across a panel of primate genomic DNA. The samples are (H) human (Homo sapiens); (C) chimpanzee (Pan troglodytes); (B) bonobo (Pan paniscus); (G) gorilla (Gorilla gorilla); (O) orangutan (Pongo pygmaeus); (R) rhesus macaque (Macaca mulatta); (P) pigtailed macaque (Macaca nemestrina); (W) common woolly monkey (Lagothrix lagotricha); (S) black-handed spider monkey (Ateles geoffroyi); (T) red-chested mustached tamarin (Saguinus labiatus); (L) ring-tailed lemur (Lemur catta); and (−C) the negative control. Vertical lines distinguish the apes, Old World monkeys (the macaques), New World monkeys, and the lemur (a prosimian); see Supplemental Figure S1 for a simple phylogeny of the primates using the same letter codes. Unusual PCR products are numbered as follows: (1) a non-ZNF sequence; (2) LLNL745 (a pseudogene whose closest paralog is ZNF100); (3) a LLNL745-like sequence in gorilla. The + indicates a weak band that was confirmed to be present. Primers were designed from the known genomic sequences and therefore may not identically match all species, resulting in variation in PCR efficiency. LLNL745 may have been deleted in chimpanzee (Newman et al. 2005) and is not in the current chimpanzee genome assembly. (B) Zinc finger alignment hypothesis for the ZNF431 clade. The alignment includes the zinc finger array of the predicted protein for selected human paralogs (H) and their respective orthologs (if found) in chimpanzee (C) and rhesus (R) genomic data. Each box represents a zinc finger; gaps are added when one locus has added or deleted a zinc finger so that flanking fingers remain aligned. The amino acid codes inside each box (for positions −1, 3, and 6, relative to the start of the α-helix) are variable positions involved in sequence-specific DNA target recognition and binding. Degenerate fingers are shaded. Frameshifts in the genomic sequences are indicated by arrows under the fingers, but for the nonhuman primate sequences these are “repaired” to match the orthologous sequences due to the incomplete nature of the nonhuman sequence data. ZNF493 and ZNF738 are not shown due to high-sequence divergence and difficulty in alignment with the other members of the clade; these genes are depicted in Figure 5.











