Segmental duplication detected in vertebrate genome sequence assemblies
Click on table to view larger version.
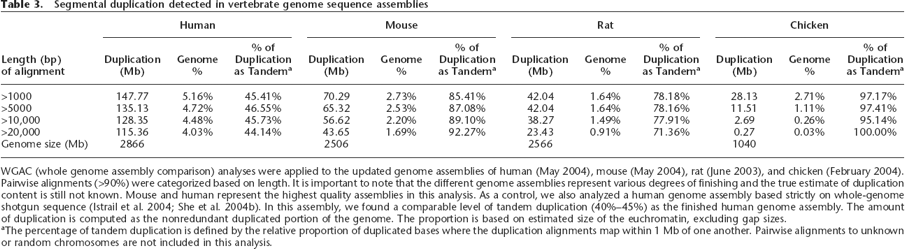
WGAC (whole genome assembly comparison) analyses were applied to the updated genome assemblies of human (May 2004), mouse (May 2004), rat (June 2003), and chicken (February 2004). Pairwise alignments (>90%) were categorized based on length. It is important to note that the different genome assemblies represent various degrees of finishing and the true estimate of duplication content is still not known. Mouse and human represent the highest quality assemblies in this analysis. As a control, we also analyzed a human genome assembly based strictly on whole-genome shotgun sequence (Istrail et al. 2004; She et al. 2004b). In this assembly, we found a comparable level of tandem duplication (40%–45%) as the finished human genome assembly. The amount of duplication is computed as the nonredundant duplicated portion of the genome. The proportion is based on estimated size of the euchromatin, excluding gap sizes.
aThe percentage of tandem duplication is defined by the relative proportion of duplicated bases where the duplication alignments map within 1 Mb of one another. Pairwise alignments to unknown or random chromosomes are not included in this analysis.











