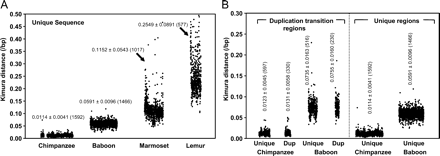
Single nucleotide substitution in unique and duplicated genomic regions. (A) Divergence of unique genomic regions. A scatter plot of genetic distances (changes/basepair, Kimura two-parameter model) determined from nonoverlapping 3-kb sliding windows for human–chimpanzee (5.0 Mb), human–baboon (5.0 Mb), human–marmoset (4.0 Mb), and human–lemur (2.8 Mb) sequence alignments. A total of 56 marmoset windows and 182 lemur windows were >0.50 in Kimura distance and thus not shown. The mean ± standard deviation (from the number of windows) are shown for each comparison based on the number of windows assessed (see Supplemental Table S1a for more details). (B) Divergence of duplicated genomic regions. Genomic sequence alignments that contain segmental duplications and that transition into unique sequence were examined for divergence between humans and nonhuman primates (baboon and chimpanzee). Duplicated and unique portions were considered separately in this analysis and compared with unique regions in A. Duplicated regions show an increase in substitution irrespective of their duplication or unique status based on alignment to the human genome.











