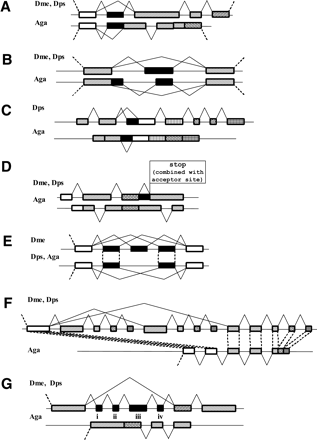
Schematic representation of the exon–intron structure and alternative splicing of the D. melanogaster genes and their orthologs in D. pseudoobscura and A. gambiae. Gene names and a brief description of events are given; for a more detailed discussion see the text. Homologous regions are shown by similar shading patterns. (A) CG1517: A cassette exon in the drosophilas corresponds to an alternative acceptor site in anopheles; (B) CG31536: A cassette exon in the drosophilas corresponds to a combination of a cassette exon and alternative donor site in the anopheles gene; (C) GC1587: An alternative acceptor site in the drosophilas corresponds to a candidate retained intron in the anopheles gene (the splicing sites are conserved in a retroposed gene); (D) GC1968: Retained intron in drosophilas, missing in the anopheles gene; numerous intron losses/insertions; (E) GC31116: Cassette exons in drosohilas that are missing in the anopheles gene; (F) GC30427: Duplication of a chain of cassette exons in drosophilas; (G) 14-3-3zeta: An additional mutually exclusive exon in Dme.











