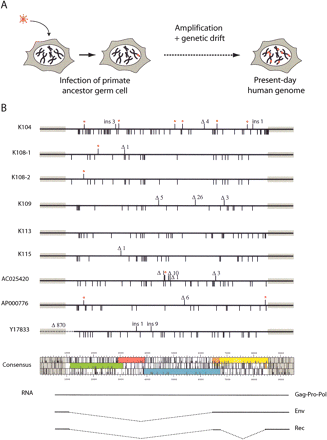
HERV-K(HML2) “endogenization” and present-day human proviruses. (A) Evolutionary scheme for HERV-K(HML2) entry into and invasion of the genome of primates. (B) Map of the full-length 9.4-kb long human-specific HERV-K(HML2) proviruses and comparison with the in silico-engineered consensus sequence. Each provirus is represented by a solid dark line, with the amino acid substitutions in Gag, Pro, Pol, and Env as compared with the consensus element indicated below the line, and the insertions/deletions (ins/Δ) and premature Stop codons (red stars) indicated above the line. The ORF map of the consensus provirus is shown, with gag in green, pro in pink, pol in blue, env in orange and yellow, the bipartite rec in orange, and the two LTRs as gray boxes. (Note that the first coding exon of rec belongs to the env ORF). The transcripts responsible for the expression of the viral proteins, with the corresponding spliced out domains (dotted lines), are schematized below the ORF map.











