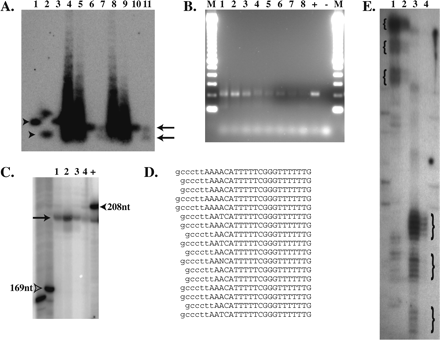
Evidence for Nucleosome Positioning on psx1. (A) Southern blot probed for psx1. Arrowheads indicate uncut (lanes 1,3) and ApoI cut (lane 2) cloned control psx1 DNA, while arrows indicate positions of uncut (lanes 6,10) and ApoI cut (lanes 7,11) mononucleosome core bands. Lanes 4, 5, 8, and 9 contain nucleosome ladders from micrococcal nuclease-digested chromatin. (B) Agarose gel electrophoresis of unique PCR product bands indicating positioning of micrococcal nuclease termini on psx1. Lanes 1–8 correspond to PCR products from reactions using ligated cores isolated with conditions 1, 3, 4, 5, 6, 7, 8, and 9, respectively (Table 1). “M”, “+”, and “−” indicate lanes containing 100-bp ladder markers (Invitrogen), positive control PCR reaction, and negative control (no template) PCR reactions, respectively. (C) Polyacrylamide gel electrophoresis with lanes 1–4 and + containing end-labeled PCR products from corresponding reactions in lanes 1–4 and + in B. Arrow indicates uniform size of psx1 core products, while open arrowhead indicates 169-nt marker and filled arrowhead indicates control reaction product of 208 nt (∼6 nt larger than the psx1 core products). (D) Sequences of vector-core junctions of psx1 nucleosome cores cloned from PCR reactions shown in lanes 1 and 2 of B. Sequence from the cloning vector is indicated in lowercase, and sequence from psx1 cores is indicated in uppercase letters, respectively. (E) Polyacrylamide gel electrophoresis of psx1 primer extension products from primer AF-SJ-6 with 10 rounds (lane 1) or one round (lane 2) of extension, and from primer AF-SJ-7 with 10 rounds (lane 3) or one round (lane 4) of extension. Curly braces demark positions of three different size ranges of products seen in both sets of primers with the largest size products in each sample being the most abundant as observed in the lanes in which samples underwent only a single round of extension, 2 and 4.











