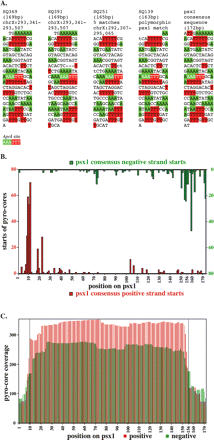
A reiterated genetic element with constrained core positioning. (A) Sequences of four closely matching cores from the original (manually isolated) series of 346 clones. These four clones, isolated from three different chromatin digestion conditions and four separate ligations, correspond to variants of a repeated sequence, designated psx1, present in the C. elegans genome (Bao and Eddy 2002). Individual polymorphisms relative to a consensus version (right) of psx1 are printed in lowercase. Runs of two or more consecutive A’s or T’s are highlighted in green and red, respectively and the sequences are displayed in 10-nt sections. (B) Starts of pyro-core sequences mapping to psx1 copies. Red and green are data from positive and negative starts, respectively. (C) Coverage plot of positive (red) and negative (green) pyro-core data for psx1.











