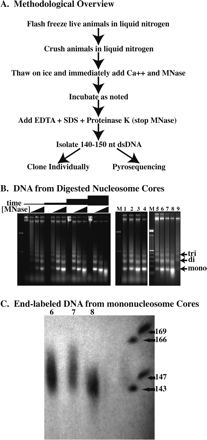
Digestion of C. elegans chromatin in situ to generate core fragments. (A) Overview of the methodology used to isolate nucleosome core DNAs. (B) Electorphoretic gel analysis of micrococcal nuclease-digested chromatin showing the descending-ladder effect with increased micrococcal nuclease digestion. Gel at left shows digestions performed with increasing concentrations of micrococcal nuclease (0, 0.2, 0.8, and 3.2 U/μL) for various amounts of time (2, 6, 18, and 54 min) at 25°C. Lanes 1–9 of the right gel correspond to DNA isolated from the various digestion conditions (digests #1–#9) as described in Table 1. (C) Sequencing gel showing end-labeled mononucleosome core DNA from digestion conditions #6, #7, and #8 in lanes 6, 7, and 8, respectively.











