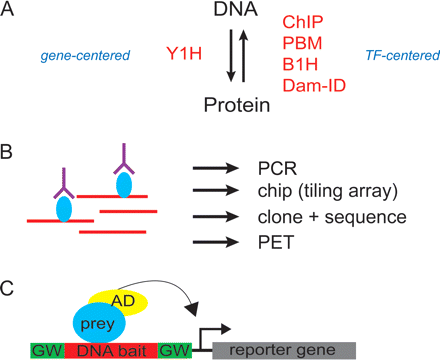
High-throughput methods for protein–DNA interaction mapping. (A) protein–DNA interactions can be mapped using either TF- or gene-centered methods, as indicated by the arrows. Y1H, yeast one-hybrid assays; ChIP, chromatin-immunoprecipitations; PBM, protein binding microarray; B1H, bacterial one-hybrid system; Dam-ID, DNA adenine methytransferase-ID. (B) ChIP is the most commonly used TF-centered method. It is based on the precipitation of a TF (blue) and its associated DNA fragments (red) using an anti-TF antibody (purple). Multiple readouts of the precipitated DNA can be used, including PCR with specific primers, tiling microarrays (chip), cloning and sequencing, and paired-end ditag sequencing. (C) Y1H assays are based on interactions of hybrid “prey” proteins with a DNA “bait” of interest. The hybrid protein consists of a protein that can bind DNA (blue) and a heterologous transcription activation domain (AD, yellow). The use of such a domain enables the identification of both activators and repressors of transcription. The readout for a protein–DNA interaction is the expression of one or more reporter genes. Prey identity is determined by PCR and sequencing. In high-throughput Y1H assays, vectors containing Gateway recombination sites (GW) are used to enable standardized cloning from promoterome resources.











