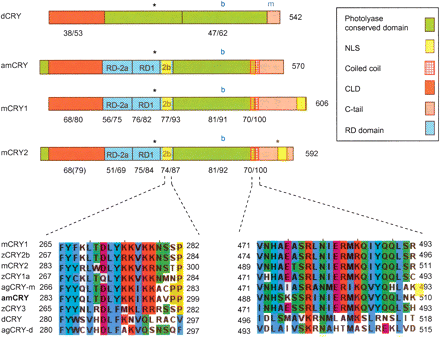
Schematic presentation of putative functional domains and motifs in Cryptochrome proteins from mouse (mCRY1 and mCRY2), honey bee (amCRY), and fruit fly (dCRY). See legend for domain/motif identity and text for additional details on each domain. Phosphorylation sites for MAPK (Sanada et al. 2004) are marked with asterisks. The blue “b” and “m” mark the location of cryb (D401N mutation) and crym (truncation of the last 19 residues in Drosophila) mutations in dCRY. Low panels show multiple sequence alignments of a putative nuclear localization signal (NLS) in the RD2b domain (left) and the coiled-coil region (right). Aligned are mammalian-type (mouse, mCRY1 and mCRY2; Zebrafise, zCRY1–3; mosquito, agCRY-m; honey bee, amCRY) and Drosophila-type (Drosophila, dCRY; mosquito, agCRY-d) CRY proteins. zCRY3 is a mammalian-type CRY protein but has no transcription repressing function in vitro according to the method of Hirayama et al. (2003).











