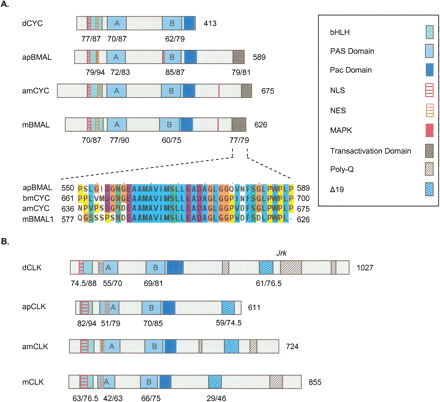
Schematic presentation of putative functional domains and motifs on Cycle/Bmal and Clock proteins from the honey bee Apis mellifera, the fruit fly Drosophila melanogaster, the giant silk moth Anthereae pernyi, and the mouse Mus musculus. See legend for domain and motif identity and text for additional details on each domain. Numbers below domains indicate identity/similarity with corresponding sequences on the honey bee ortholog. The numbers at the end of each diagram indicate protein size (number of amino acid residues). (A) Cycle/Bmal proteins. Inset shows a CLUSTALW multiple sequence alignment of a putative domain that is thought to be necessary for apBMAL and mBMAL transcriptional activity (see text for details). bmCYC is the CYC ortholog from the moth Bombyx mori. Alignments were generated with CLUSTALW and colored with JalView according to the default CLUSTALX convention. (B) Clock proteins.











