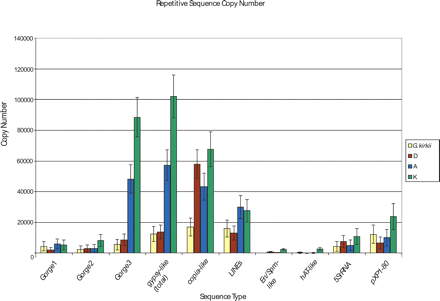
Copy number estimates for repetitive sequences in Gossypium. Copy numbers for repetitive sequences recovered in the WGS libraries were estimated as described (see Methods). The majority of repetitive sequences are LTR retrotransposons, particularly in the larger-genome species. In both the A and K genomes, massive amplification of Gorge3 gypsy-like sequences has occurred, contributing predominantly to genome size expansion in these two lineages. In the smallest Gossypium genome, G. raimondii (D genome), and copia-like sequences have proliferated and are primarily responsible for genome size expansion in this lineage. Class II sequences were less abundant and appear to contribute little to genome size evolution in the genus. Tandem repeats are approximately evenly distributed among all four species, with pXP1–80 sequences slightly elevated in G. exiguum (K genome).











