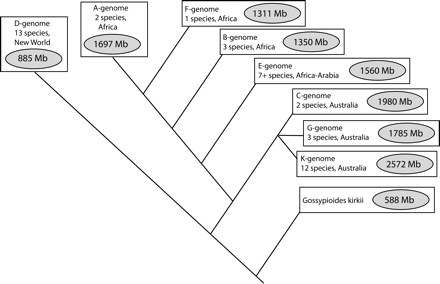
Evolutionary relationships among diploid members of Gossypium. Gossypium is a monophyletic genus composed of ∼50 species that are widely distributed throughout many tropical and subtropical regions. Diploid species have a haploid complement of 13 chromosomes. Gossypium is divided into eight genome groups based on cytogenetic data and level of fertility in interspecific hybrids (Endrizzi et al. 1985). Multiple molecular data sets support the phylogenetic relationships indicated, including the outgroup relationship of Gossypioides kirkii (Wendel and Albert 1992; Seelanan et al. 1997; Small et al. 1998, 1999). Despite conservation of chromosome number among the diploids, genome size varies threefold, from an average of 885 Mb in the New World D-genome species to an average of 2576 Mb in the Australian K-genome species (Hendrix and Stewart 2005).











