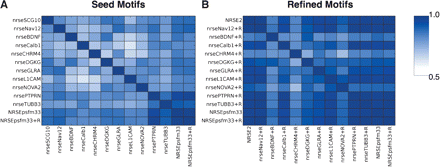
Different seed motifs converge following motif refinement. (A) A total of 10 initial seed motifs from known or predicted sites are compared using the motif similarity score (see Methods) to our starting motif (SCG10) as well as a PSFM of 33 known instances (NRSEpsfm33) and its refined version (NRSEpsfm33+R). The correlations median is 0.80. (B) Motif refinement of SCG10 (called NRSE2) and of the 10 initial motifs (denoted with a +R) are markedly more similar, with a motif correlations median of 0.91 with several intermotif correlations rising above 0.95.











