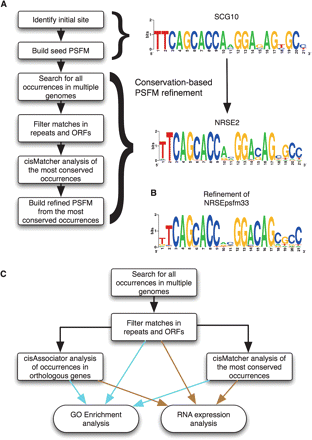
Experimental approach. (A) Results from genome-wide matches to the initial NRSE PSFM (SCG10) were analyzed with cisMatcher and used to create a refined NRSE PSFM (NRSE2). (B) A refinement starting with a PSFM of 33 known sites (Supplemental Table S2) produces a result very similar to NRSE2. (C) The genome was searched for occur- rences of NRSE1, using its consensus (TYAGMRCCNNRGMSAG) (Bruce et al. 2004); or NRSE2, using its position-specific frequency matrix (PSFM). Resulting NRSE1 and NRSE2 gene cohorts were then analyzed for Gene Ontology (GO) enrichment and expression analysis as follows: (1) the NRSE2 PSFM was further processed and analyzed for GO enrichment and expression analysis of two subsets; (2) human genes with matches that co-occur in mouse and/or dog, and (3) human genes that are nearest to the “most conserved” matches, as identified by cisMatcher.











