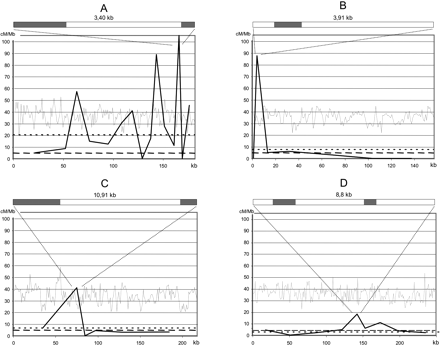
Fine-scale analysis of the distribution of CO breakpoints in four intervals. (A) interval 70: 44.5kb-3; 16.4kb-2; 6.2kb-5; 19.5kb-4; 17.5kb-3; 9.4kb-4; 12.3kb-7; 8.1kb-0; 8.3kb-2; 3.2kb-4; 13.0kb-5; 6.4kb-1; 3.4kb-5; 3.3kb-0; 11.0kb-7. (B) interval 21: 2.5kb-0; 3.9kb-5; 13.8kb-1; 23.6kb-2; 45.7kb-2; 29.2kb-0; 38.6kb-0. (C) interval 57: 69.5kb-6; 12.2kb-7; 7.1kb-0; 17.4kb-1; 46.5kb-2; 65.0kb-3. (D) interval 37: 35.2kb-2; 32.7kb-0; 38.1kb-1; 8.8kb-2; 12.6kb-1; 28.3kb-1; 20.2kb-1; 49.2kb-2; 38.7kb-1. For each interval the size of each DNA fragment is given and the number of recombinant plants. (Light gray line) G+C content calculated in 1-kb windows. (Black line) CO rates in centiMorgans per megabase. (Small dotted line) interval COs rate average. (Large dotted line) chromosome 4 COs rate average. Above each major peak is the gene organization of the fragment. (Gray box) gene.











