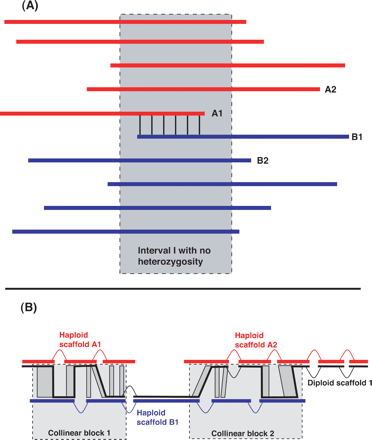
The splitting rule (A) and a diploid scaffold (B). (A) Suppose that the red reads are of one haplotype and the blue reads are of the other haplotype, and that all reads have computationally detected overlaps with all other reads of the same color. In principle, the red reads should assemble in one contig, and the blue reads in another contig. However, if A1 and B1 overlap only in a region of haplotype identity, then A1 and B1 will also have a computationally detected overlap. When Arachne builds contigs, this overlap would trigger algorithms designed to prevent assembly through repeats and break both the red and blue scaffolds. The splitting rule was devised to recognize this situation and sever the overlap between A1 and B1 prior to the formation of contigs, thus allowing the red and blue contigs to assemble separately. See text for details. (B) In general, a diploid scaffold is an alternation between regions represented by a single haploid scaffold and regions represented by a collinear block of alignments relating two haploid scaffolds. For example, the diploid scaffold in B has two collinear blocks and three regions represented by a single haplotype. Within each collinear block, the reference path (thick black line) is chosen to minimize the number of contig gaps.











