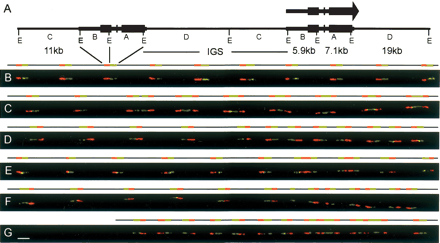
Structural analysis of the human rDNA locus by molecular combing. (A) Schematic representation of two canonical rDNA units. Restriction with EcoRI (sites E) yields four distinct fragments spanning the transcribed region (thick line) and the intergenic spacer or IGS (thin line). The orientation of the rRNA transcript and the positions of the ribosomal genes (black boxes) are shown. (B) Two-color hybridization on combed human DNA. The red probe is the 5′ EcoRI fragment B detected with Texas Red. The green probe is the 3′ fragment A detected with FITC. The image displays 10 canonical rDNA units in tandem, each composed of a dual fluorescent signal and the adjacent nonhybridizing spacer segments. (C) Hybridizations of the probes on human DNA that illustrates noncanonical units. The image displays a region containing two canonical units (left) followed by seven palindromic units, with each half joined by its 3′ region (3′–3′ palindromes) and separated by short IGS segments. (D) Hybridizations illustrating successive inverted units with gaps. The image displays five canonical units, followed by a series of seven palindromic units separated by short gaps. (E) Hybridizations illustrating 5′–5′ palindromic units. The image displays six canonical units (left), followed by two gapless palindromic units joined at their 5′ extremities and two canonical units (separated by a short IGS) in an inverted orientation with respect to the canonical units on the left. (F) Hybridizations illustrating 5′–5′ palindromic units with gaps. The image displays six canonical units, followed by a single 3′ fragment and three 5′–5′ palindromes with short central gaps separated by short IGS segments. (G) Hybridization illustrating complex recombinant structures. The image displays closely spaced, inverted units (left), followed by two complex units with alternating 5′ and 3′ coding sequences, separated by short IGS sequences, and one canonical unit (right). Scale bar, 10 kb, applicable to all images.











