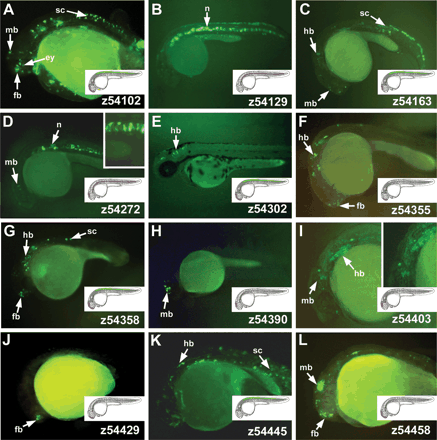
Highly conserved non-coding regions (HCNRs) located between Irx3 and Irx5 genes show enhancer activity in zebrafish. Lateral views of 24-hpf zebrafish showing enhanced green fluorescent protein localization promoted by different HCNRs (A–L). Insets show magnifications of some of the corresponding expression domains. The number in the lower righthand corner of each panel corresponds to the position of the human homologous region in chromosome 16 (NCBI build 33) as shown in Figure 1I. For better comparison, illustrations show the expression domains in which the HCNRs are active. Color codes are as those in Figure 1. Note that this is an oversimplification scheme, as many enhancers are active in subdomains of these territories. (fb) Forebrain; (mb) midbrain; (hb) hindbrain; (sp) spinal cord; (ey) eye; (n) notochord.











