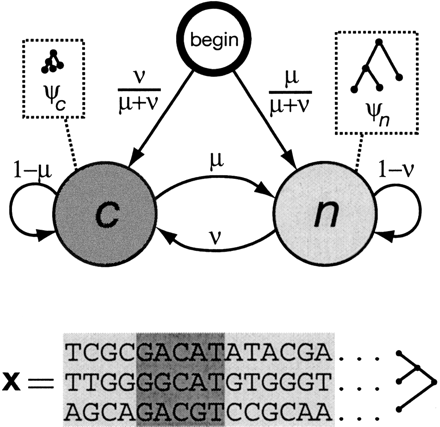
State-transition diagram for the phylo-HMM used by phastCons, which consists of a state for conserved regions (c) and a state for nonconserved regions (n). Each state is associated with a phylogenetic model (ψc and ψn); these models are identical except for a scaling parameter ρ (0 ≤ ρ ≤ 1), which is applied to the branch lengths of ψc and represents the average rate of substitution in conserved regions as a fraction of the average rate in nonconserved regions (see Methods). Two parameters, μ and ν (0 ≤ μ, ν ≤ 1), define all state-transition probabilities, as illustrated. The probability of visiting each state first (indicated by arcs from the node labeled “begin”) is simply set equal to the probability of that state at equilibrium (stationarity). The model can be thought of as a probabilistic machine that “generates” a multiple alignment, consisting of alternating sequences of conserved (dark gray) and nonconserved (light gray) alignment columns (see example at bottom).











