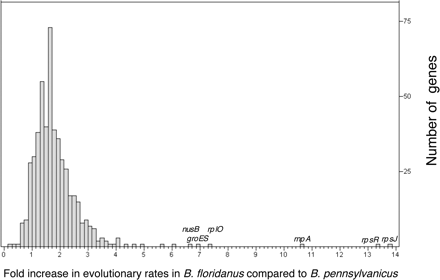
Accelerated rates of evolution in the lineage leading to B. floridanus compared to B. pennsylvanicus, since their divergence from a common ancestor. Rates of protein divergence were compared using a relative rates test with E. coli as an outgroup (see Methods). The analysis included 516 proteins for which RSD identified orthologs in the two genomes, and for which divergences between the two Blochmannia or between either endosymbiont and E. coli did not exceed 2.0. On average, proteins evolved 1.88 times faster in B. floridanus compared to in B. pennsylvanicus. Only 50 of the proteins tested evolved more slowly in B. pennsylvanicus than in B. floridanus or at the same rate, with values ≤1on the histogram, while 467 genes evolved faster in B. floridanus. Supplemental Table S4 lists proteins with particularly accelerated evolutionary rates in B. floridanus.











