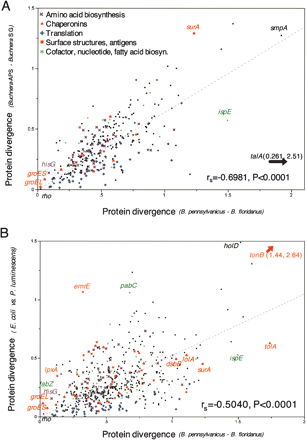
Correlated protein divergences suggest parallel selective constraints in endosymbionts and free-living bacteria. Protein divergence in Blochmannia shows a strong correlation with divergence at orthologous genes of (A) Buchnera (–A. pisum versus –S. graminum) and (B) E. coli versus P. luminescens. Lines that best fit the data and intercept zero are shown, but significance was tested using the nonparametric test of association (JMP 4.0; SAS Institute Inc.). For all bacterial pairs, the most conserved genes include certain chaperonins and translation functions (mostly ribosomal proteins), while the most divergent genes include surface structures. Substitution rates above 2 are prone to saturation but are included here for comparison. Supplemental Table S3 lists all of the pairwise divergences.











