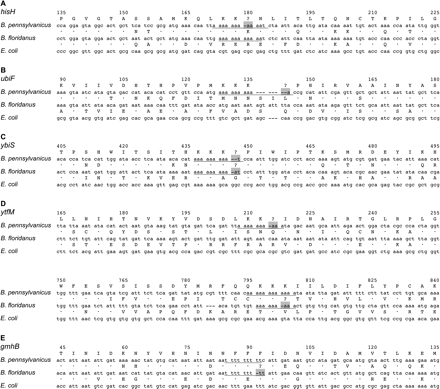
Sequence context of homopolymeric frameshift mutations within five Blochmannia genes. The single frameshifts in Blochmannia genes (A) hisH, (B) ubiF, (C) ybiS, (D) ytfM, and (E) gmhB were “corrected” manually, based on comparisons with sequences that lack the frameshift. Genes were then aligned by their inferred amino acid sequence against E. coli. Underlined nucleotides indicate poly(A) and poly(T) tracts in which frameshifts occur. Regions highlighted in gray indicate the putative sites of transcriptional or translational slippage that may restore the proper reading frame. (•) Amino acid identical to the top sequence (B. pennsylvanicus).











