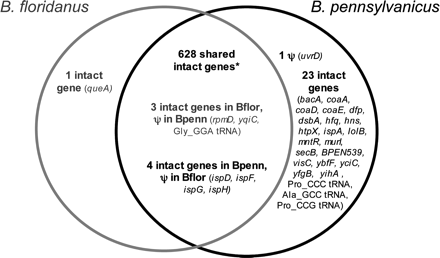
Comparison of B. pennsylvanicus and B. floridanus gene contents. Genes in the outer section of each circle are unique to one genome, while those listed in the intersection of the two circles are shared. Note that seven shared genes are apparently functional in one genome, but pseudogenes in the other. The truncation of B. floridanus dnaX (see text) is not included as a pseudogene here. The fusion of yidCD in B. pennsylvanicus is counted as two intact genes. (*) The 628 shared intact genes include genes that have a single frameshift within long poly(A) tracts but are otherwise intact (see text). These loci include B. pennsylvanicus hisH, ytfM, ubiF, and ybiS and B. floridanus ytfM, ybiS, and gmhB.











