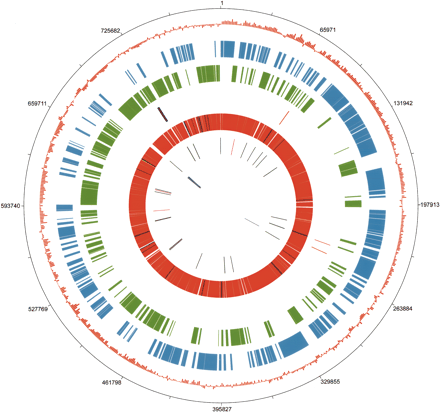
Circular map of the B. pennsylvanicus genome and genome features. The origin of replication was putatively set upstream of gidA, in accordance with the shift of GC skew. The concentric rings denote the following features: (1) numbered base pair coordinates beginning with base one of the gidA open reading frame (ORF); (2) GC skew as calculated by (G–C)/(G+C) using a 2.5-kb sliding window; (3) ORFS present on the leading strand (+); (4) ORFS present on the lagging strand (–); (5) pseudogenes of B. pennsylvanicus that are present in (orange) or absent from (black) B. floridanus; (6) ORFS of B. pennsylvanicus that are shared with (red), pseudogenes in (yellow), or absent in (dark blue) the genome of B. floridanus; (7) transfer RNAs (tRNAs) that are shared with (gray) or absent in (pink) B. floridanus; (8) ribosomal RNAs (rRNAs) 5S, 16S, and 23S (purple). Circular plot and GC skew analysis were generated by the JENA Prokaryotic Genome Viewer–Export Version v3.06.04.











