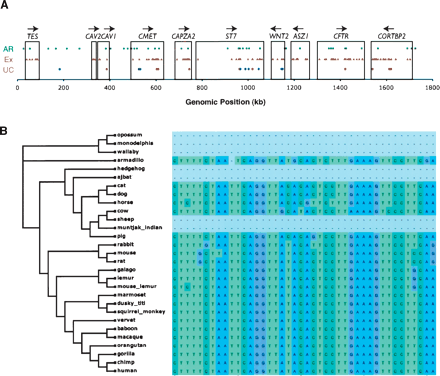
Ultraconserved and mobile-element derived constrained elements. (A) The locations of three types of features are shown along the genomic coordinates of the human locus as follows: green squares indicate the locations of ancestral repeats that overlap constrained elements; orange triangles correspond to exons; and circles correspond to the ultraconserved elements (Table 2), broken down according to those that overlap exons (red), and those that do not (blue). RefSeq genes and their transcriptional orientation are marked by boxes and arrows, respectively. (B) A small alignment region corresponding to an ancestral repeat region (part of an L3 element) overlapping a constrained element scoring >100 rejected substitutions. Nucleotides are color-coded, and gaps are indicated in gray (the fully gapped placental species are missing data). The displayed region corresponds to positions 495,140–495,181 of the human sequence (with the first base of the locus being position 1).











