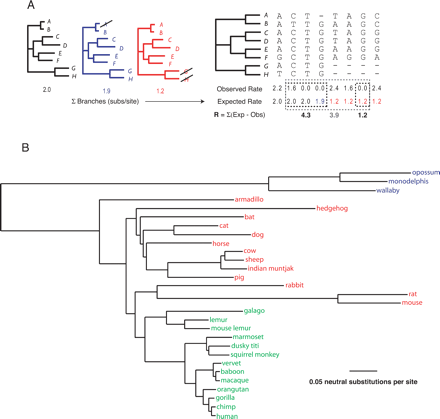
Overview of GERP. (A) Each column of the compressed alignment (corresponding to each base of the human sequence) is analyzed independently. Number of substitution events is inferred, giving “observed” values (see Methods); the “expected” rate for each column is determined by summing the branches of the neutral tree that remain after removing species with a gap character (compare the black, red, and blue neutral trees with the correspondingly colored expected rates). Candidate constrained regions are identified as consecutive columns of observed rates smaller than the expected rates (black boxes). Nearby candidates are merged (gray box) across a limited number of unconstrained columns. Finally, each candidate is scored as the sum of the deviations from expectation at each column, collectively termed as “rejected substitutions.” (B) Neutral tree for the complete set of species analyzed here (see Methods); the tree is rooted arbitrarily for display purposes only, and analyses are performed using an unrooted tree. Primates are in green, non-primate placental mammals are in red, and marsupials are in blue.











