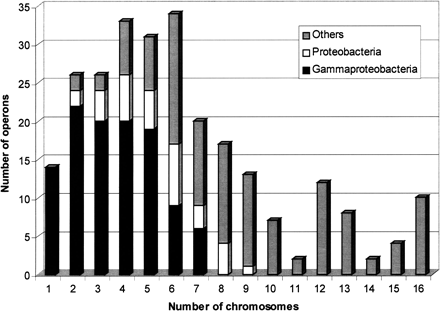
Diagram of the phylogenetic distribution of 245 E. coli operons (of 309) fully recovered by at least one domain team in the set of 15 Gram-negative bacteria. The figure shows the distribution of the operons as a function of the number of chromosomes in which the operons were identified as syntenic. Each class has been divided into three categories, depending on the species where the teams were found, i.e., only in gammaproteobacteria or only in proteobacteria, or also in other taxons. Thus 96 operons (gray) were recovered only within close species (gammaproteobacteria), but the diagram shows that 149 other operons are conserved in more distant bacteria. Fourteen operons (class 1) were found only in E. coli.











