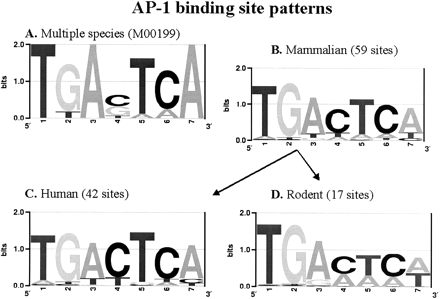
AP-1 binding site preferences in different species clusters. The LOGOs for the AP-1 binding preferences from (A) TRANSFAC weight matrix M00199 (includes sites from various organisms like human, mouse, rat, chicken and frog) and (B) human, mouse, and rat. Most of current algorithms are using one of the two variants of weight matrices to scan the promoter regions. (C, D) Further partition of the mammalian sequences into human and rodent, revealing differences in the suboptimal patterns. LOGOs are created by using the program enoLOGOS (Workman et al. 2005) available on the Web http://biodev.hgen.pitt.edu/cgi-bin/enologos/enologos.cgi.











