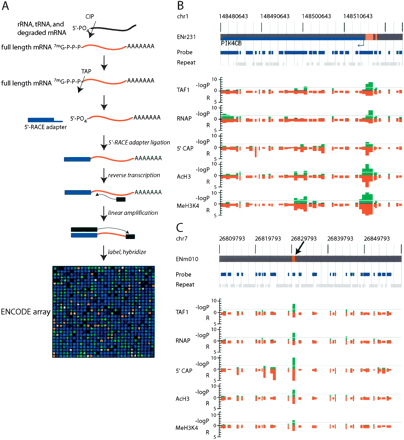
Experimental validation of the putative promoters by detection of 5′-capped mRNA. (A) A schematic of ARLM-RACE used to map 5′-capped mRNA sequences within the ENCODE regions. (CIP) Calf Intestine Alkaline Phosphatase; (TAP) Tobacco Acid Pyrophosphatase (the decapping enzyme). A detailed procedure is described in the Methods. (B) Detection of the 5′-capped mRNA sequence from and localized acetylated histone H3 (AcH3) and methylated histone H3 lysine residue 4 (MeH3K4) at a known promoter (marked with yellow box). Negative logarithmic P values of enrichment by RNAP ChIP, TAF250 ChIP, ARLM-RACE, Acetyl Histone H3, and Methyl K4 Histone H3 for each probe are plotted in green and relative enrichment (R) values for each probe fragment are plotted in red on the inverted axis. (C) Detection of the 5′-capped mRNA sequence from a previously unknown promoter (marked with magenta box and denoted by an arrow). Negative logarithmic P values of enrichment by RNAP ChIP, TAF250 ChIP, ARLM-RACE, Acetyl Histone H3, and Methyl K4 Histone H3 for each probe are plotted in green and relative enrichment (R) values for each probe fragment are plotted in red on the inverted axis.











