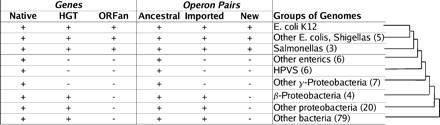
The evolutionary history of genes and operons. For each gene in E. coli K12, we determined which groups of genomes contained a potential ortholog of that gene and classified genes as native, HGT, or ORFan. We performed a similar analysis on each adjacent pair of genes predicted to be in the same operon and classified pairs as ancestral, imported, or new. Some genes and pairs could not be classified. We show examples of patterns of presence or absence for each class of gene and for each class of operon pair. The placement of the genomes at varying distances from E. coli K12 is in accordance with generally accepted phylogenies and with a whole-genome protein sequence tree (P. Dehal and E.J. Alm, unpubl.). “Other enterics” includes Yersinia, Buchnera, and Wigglesworthia species; “HPVS” includes Haemophilus, Pasteurella, Vibrio, and Shewanella species; and “other γ-Proteobacteria” includes Pseudomonas, Xanthomonas, and Xylella species. For the inferred histories to be correct, the union of all groups up to a given age must be monophyletic, but each outgroup need not be. For example, we believe that HPVS and the Enterobacteria together form a monophyletic clade but not HPVS by themselves.











