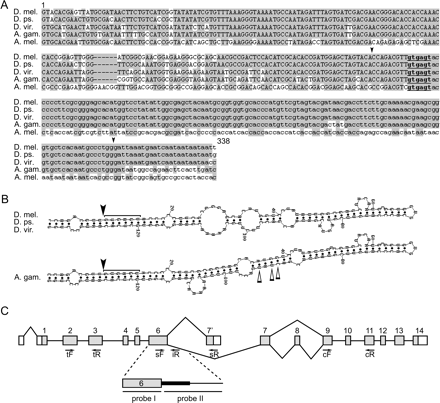
(A) Alignment of five orthologous fragments of the hth gene from various insects. The species are Drosophila melanogaster (D. mel.), Drosophila pseudoobscura (D. ps.), Drosophila virilis (D. vir.), Anopheles gambiae (A. gam.), and Apis melifera (A. mel.). The conserved intronic sequence is shown in lowercase; uppercase represents the adjacent constitutive exon 6. The donor splice-site consensus is underlined. Black arrowheads show the start and end of the predicted hairpin fold region in the first four species.(B) RNA hairpin structures of the highly conserved intronic sequence predicted by MFOLD 3.1. The donor splice-site consensus is marked by the bracket. Splice-site positions are indicated by black arrowheads. Open arrows show substituted nucleotides in the A. gambiae sequence. (C) Splicing diagram of the hth gene (not shown to scale). Gray and white boxes represent protein-coding and noncoding exons, respectively. Straight lines connect constitutively spliced exons. Angled lines show sites of alternative splicing. The transcript BT010238 terminates at the exon 7′. Positions of primers used in real-time PCR are indicated by arrows. The fragment amplified by tF/tR (HTH-t) primer pair is present in all hth RNA transcripts, and thus, detects total transcriptional output of the hth gene. The fragment amplified by sF/sR (HTH-s) primer pair detects the presence of spliced hth mRNA (GenBank accession BT010238). The fragment amplified by sF/iR (HTH-i) primer pair spans the border of the constitutive exon and the adjacent intron, and thus detects the presence of intron-retained hth RNA isoforms. A fragment amplified by cF/cR (HTH-c) primer pair is present in all hth RNA transcripts except BT010238, and thus, detects abundance of the remaining hth transcripts. The enlarged portion of the diagram shows the position of the ultraconserved element (thick black line) and the location of DNA probes used in RNA-blot hybridization.











