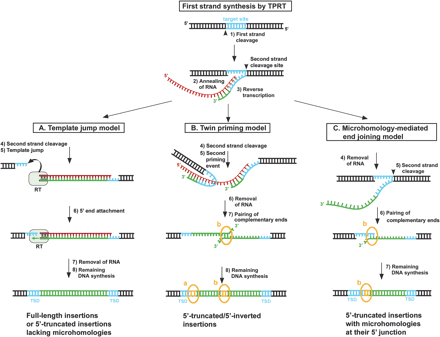
Alternative models for the attachment of the 3′-end of the L1 cDNA to the chromosomal target DNA. According to the TPRT model, L1-encoded EN nicks the bottom strand of the target DNA (1) and exposes a 3′-hydroxyl that primes reverse transcription of the element's full-length RNA (2, 3). (A) Template-jump model: After first-strand synthesis is completed, second-strand cleavage occurs (4), and the L1 RT jumps from the L1 RNA template onto the upstream target DNA (5). cDNA synthesis is continued using the genomic 3′ overhang as template, and the 3′-end of the cDNA is attached to the genomic DNA (6). Jumps from the 5′-end of full-length RNAs generate full-length L1 insertions. If the reverse transcriptase jumps from 5′-degraded L1 RNAs, 5′-truncated elements are generated without the need for microhomologies (Eickbush 2002; Bibillo and Eickbush 2004). (B) Twin priming model: L1 EN cleaves the second DNA strand (4) before reverse transcription of the first-strand cDNA has been completed (3), producing an additional 3′-hydroxyl and a stretch of single-stranded DNA, which serves as an internal primer. This primer anneals at complementing nucleotides in an internal region of the L1 RNA template, and primes reverse transcription upstream from the 3′-end of the L1 RNA (5), where the initial TPRT started. After the RNA is removed from the RNA/cDNA structure (6), the single-stranded cDNAs pair at a small region of limited complementarity (7), and the remaining DNA synthesis is completed (8) (Ostertag and Kazazian Jr. 2001b). In this model, patches of microhomology are a consequence of either RNA/DNA annealing at the 5′ junction (a) or DNA/DNA base-pairing at the inversion junction (b). (C) Microhomology-mediated end-joining model: After initiation of TPRT, the reverse transcriptase falls off the RNA template before it has reached the 5′-end of the RNA (3). Second-strand cleavage (5) occurs after reverse transcription has stopped and the L1 RNA has been degraded (4). The 3′ overhang of the chromosomal target site anneals to the 3′-end of the L1 cDNA at a region of limited complementarity (6), primes second-strand synthesis (7), and leads to the formation of 5′-truncated L1 elements with microhomologies at their 5′ junctions (b). In all three models, the second DNA strand is joined to the target site by the action of a cellular ligase after completion of second-strand synthesis.











