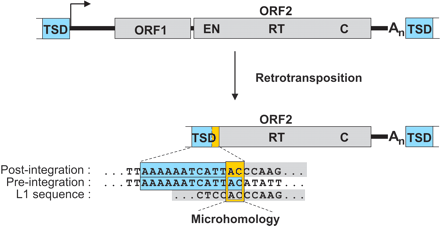
Retrotransposition of a functional human L1 element frequently results in 5′-truncated copies with overlapping nucleotides at the 5′ junctions. These microhomologies (yellow box) between the genomic target site duplication (TSD) and the 5′-end of the adjacent L1 sequence make a precise assignment of the 5′ boundary of the L1 insertion ambiguous. A 13-bp TSD sequence (blue) observed after de novo L1 retrotransposition (Symer et al. 2002) serves as a representative example. Nucleotide sequences of the genomic pre- and post-integration sites as well as the L1 consensus sequence at the junction region are shown. (ORF1/2) Open reading frame 1/2; (EN) endonuclease; (RT) reverse transcriptase; (C) cysteine-rich motif.











