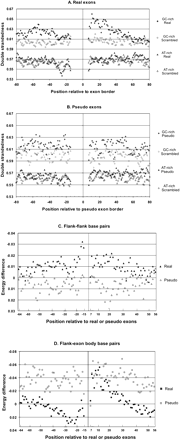
Secondary structure analysis. (A) Double strandedness in real exon flanks. Exons with their flanks were folded using Mfold. As a control, each upstream flank was scrambled and each downstream flank was scrambled, and the scrambled flanks were reconnected to the original exon body and then folded using Mfold. Both the original and scrambled versions of the sequences were divided into a HGC class (GC% of the most proximal 50-nt flanks >55) and an LGC class (GC% of the most proximal 50-nt flanks <45), leading to four different data sets as follows: original HGC (♦), original LGC (▴), scrambled HGC (⋄), and scrambled LGC (▵). We then plotted the double strandedness as a function of positions in flanks. Double strandedness reflects the frequency of all predicted base pairing at each position (see Results and Methods). (B) Double strandedness in pseudo exon flanks. Exactly the same analysis of pseudo exons as a control. (C) Flank-flank base pairing around HGC exons. The incremental contribution of each interflank base pair to the energy of each predicted stable structure was extracted from the Mfold output after folding both original exon plus flank sequences and again after scrambling the flanks. The difference between these two values (original sequence energies—scrambled sequence energies) is plotted as an indication of the excess secondary structure contributed at each position (filled symbols). A negative value represents more base pairing. For comparison, the same process was carried out on pseudo exons (open symbols). (D) Flank-exon base pairing around HGC exons. Differences in the free energy contributions of individual base pairs were calculated and displayed as in C, except only base pairs between flank and exon positions were chosen. (Real exons) Filled symbols; (pseudo exons) open symbols.











