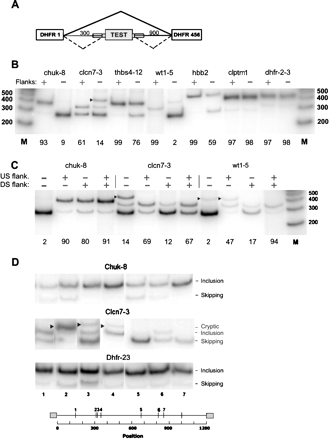
Effect of flanks on splicing. (A) Schematic diagram of the test construct used. PCR was used to insert the test exon with both, neither, or one flank of ∼50 nt beyond the splice-site consensus. (Thin lines) Introns; (large rectangles) exons; (small rectangles superimposed on lines) flanks. (B) Effect of inclusion of both flanks on splicing. Cloned plasmids were transfected into 293 cells and subjected to RT-PCR using incorporation of [32P]dATP and polyacrylamide gel electrophoresis; the intensity of the radioactive bands was quantified with a PhosphorImager. Markers are from an end-labeled 100-bp DNA ladder (Invitrogen). (chuk) Conserved helix-loop-helix ubiquitous kinase; (clcn7) chloride channel 7; (thbs4) thrombospondin 4; (wt1) Wilms tumor 1; (hbb) human β-globin; (clptm1) cleft lip and palate transmembrane protein 1; (dhfr-2-3) fused exons 2 and 3 of the Chinese hamster dhfr gene. The number after the hyphen denotes the exon number. An arrowhead indicates use of a cryptic donor splice site found by sequencing the PCR product to be gaa|gtaagt at +83 in the downstream dhfr intron 3; otherwise, the upper band position represents exon inclusion, and the lower band position represents exon skipping. The percent inclusion (radioactivity in the included band representing exon splicing divided by the total of all bands) is indicated below each panel. (C) Testing the effect of individual flanks in the case of 3 exons as in B above. (D) Splicing of the indicated flankless exons inserted at random locations within the 1200-nt intronic sequence of the test construct. In this experiment, each exon was bounded on each side by the same 25-nt bacterial sequence used for transposition. Cloned plasmids were transfected into 293 cells and analyzed by RT-PCR as in B. (Arrowheads) Splicing at unidentified cryptic sites. The seven insertion positions were at the following distances from the 5′ end of the intron: 143, 310, 321, 339, 674, 809, and 864, respectively. The insertion point for the experiments shown in B and C was 304, the natural end of intron 1 of the hamster dhfr gene.











