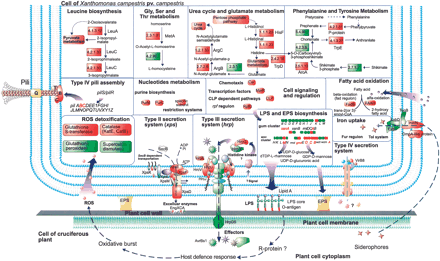
A schematic illustration of the experimentally determined interaction between Xcc and host cell. The principle pathways are shown in bold. Putative metabolic pathways were predicted by KEGG. Bacterial genes and gene products associated with pathogenicity are arranged within colored frames. Genes identified in this study are framed in red, while genes that were reported previously in xanthomonads are framed in green. (EPS) Exopolysaccharides; (FAAH) fatty acid α hydroxylase; (Hrp) hypersensitive response and pathogenicity genes; (LPS) lipopolysaccharides; (ROS) reactive oxygen species; (R-protein) resistance gene protein; (Xps) Xanthomonas protein secretion.











