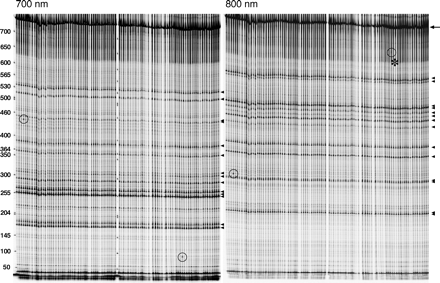
Tilling screen. Example of a gel used for mutation detection within the Sara-4 amplicon in 96 balanced mutant lines; 700 (left) and 800 nm (right) emission images showing the fragments generated by Cel-I-based heteroduplex cleavage. (Arrowheads) Fragments generated by cleavage at 14 different silent background polymorphisms in the balancer that could be reidentified by sequencing; (circles) two novel mutations. Notice that both the polymorphisms and the novel mutations appear in both channels with complementary sizes. In the case of the second mutation, the longer fragment (800-nm channel) is less distinguishable (asterisk). After sequencing, we could, however, reconfirm that this signal corresponds to a bona fide mutation. Note also that the silent polymorphisms are present in all 96 lines, reflecting the fact that the balancer is itself isogenic. New mutations appear only in individual lines. Full-length product, 747 bp (arrow). Cleavage of the two new mutants yields 440 bp (700 nm) + 315 bp (800 nm) (lane 4) and 90 bp (700 nm) and 660 bp (800 nm) (lane 80).











