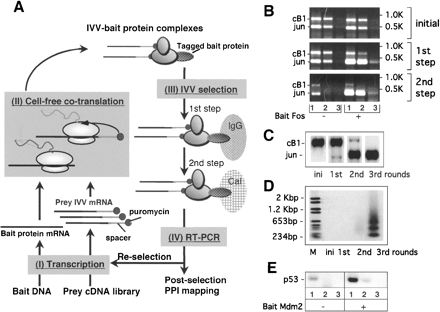
IVV selection based on cell-free cotranslation. (A) Design of the IVV selection. The selection is composed of transcription (I), cell-free cotranslation (II) coupling translation and interaction of tagged bait protein and prey IVV library to form complexes, IVV two-step selection (III) (Rigaut et al. 1999), and RT–PCR (IV) to amplify the library. (IgG) IgG beads for the protein A (z domain) tag used for the 1st step of the purification; (Cal) Calmodulin beads for the calmodulin binding protein (CBP) tag used for the 2nd step of the purification. (B) Enrichment in the 1st and the 2nd steps of the two-step selection in IVV. The enrichment was confirmed with (Fos+) or without (Fos–) 400 nM fos as bait and IVV templates of 100 nM jun and cB1 as prey. (Lanes 1–3) RT–PCR analysis with original eluate and 100-fold dilutions to 10,000-fold, respectively. (C) Enrichment after the two-step selection of IVV. RT–PCR analysis was performed with 100 nM cB1 and 10–2 nM jun. (Ini) Initial mixture of jun and cB1. (D) Enrichment of Jun from a mouse brain cDNA library. Southern blots were done with RT–PCR products of the two-step selection of IVV using Dig DNA labeling and detection kit (Roche) with a jun (616–1005) template labeled with a random hexamer primer. (Ini) Initial library. (E) Enrichment of p53 from a human brain cDNA library. PCR (1 cycle of 98°C, 2 min; 35 cycles of 98°C, 30 sec; 62°C, 30 sec; 73°C, 30 sec; one cycle of 73°C, 2 min) was done with human_p53-5′40–63 (ctgagtcaggaaa cattttcagac) and human_p53-3′257–276 (gggccaggagggggctggtg) using RT–PCR products after the 1st step of the two-step selection of IVV. The enrichment was confirmed by comparing the results with Mdm2 (+) or without Mdm2 (–) as bait. (Lanes 1–3) PCR analysis with the original eluate and 100-fold dilutions to 10,000-fold, respectively.











