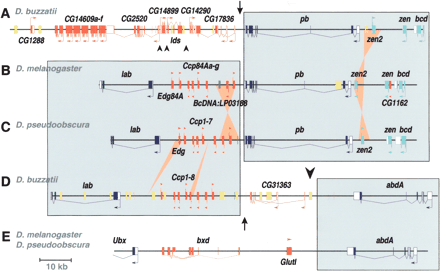
Gene organization of the lab-abdA and pb genomic regions of D. buzzatii compared with the homologous regions of D. melanogaster and D. pseudoobscura. The localization of the lab-pb split (arrow) and the Ubx-abdA split (large arrowhead) are indicated. (A) Sequence of D. buzzatii BAC 40C11 containing the pb region. (B) Organization of the lab-pb region in D. melanogaster. (C) Idem in D. pseudoobscura. (D) Sequence of D. buzzatii BAC 5H14 containing the lab-abdA region. (E) Organization of the abdA region in D. melanogaster and D. pseudoobscura. Genes are represented as open (UTRs) and filled boxes (coding sequences) with arrows indicating the sense of transcription. Hox genes are colored in dark blue, Hox-derived genes in light blue, non-Hox genes in red, noncoding RNA genes in orange, and the BcDNA:LP03188 and orthologous sequences in green. Transposable element insertions (usually ISBu elements, see Negre et al. 2003) are shown as yellow boxes. Large shaded rectangles include homologous Hox-gene regions in different species. Ochre triangles denote small inversions and insertions or deletions. Small arrowheads show breakpoints between D. buzzatii and D. melanogaster in non-Hox regions.











