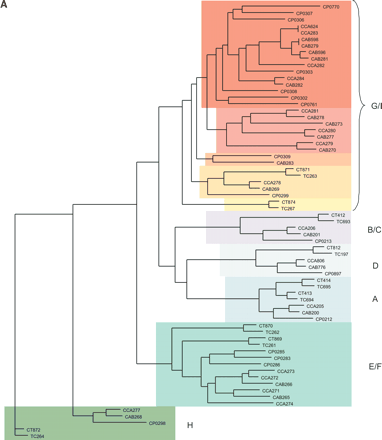
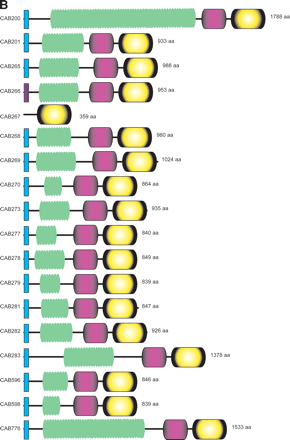
(A) Clustering of the Pmp-family proteins of sequenced chlamydial species. Phylogenetic tree of the Pmp-family proteins from Cp. abortus (S26/3), Cp. caviae (GPIC), Cp. pneumoniae (AR39), C. muridarum (Nigg), and C. trachomatis (serovar D). The Pmp protein families (A–I) are marked as previously assigned (Grimwood and Stephens 1999). (B) Architecture of the Pmp-family proteins of Cp. abortus. Conserved domains are color coded: signal sequence (light blue), transmembrane domain (dark purple), Pmp-repeat region (green), central PMP_M region (light purple), and autotransporter-like domain (yellow). The amino acid sequences of the pseudogenes CAB270 and CAB273 and the phase-variable genes CAB279 and CAB596 have been reconstructed in silico for the purpose of this analysis. CAB267 is a gene remnant.











