
Relationships between base composition, structure, and sequence divergence of large insertions of nuclear organelle DNA. (A) The G/C content of the insertion comes to resemble that of its nuclear chromosomal neighborhood. The primary insertion is
assumed to have had the same G/C content as the corresponding region of pt/mtDNA. The G/C contents of the chromosomal regions
hosting the insertions are based on those of the 300 kb of nuclear DNA immediately flanking the respective insertion. White
boxes indicate segments with an overall similarity of >99.5% to pt/mtDNA; the insertions indicated as gray boxes are less
similar to organellar DNA. The two integrants with the highest transition/transversion ratios are indicated in bold. (B) Relationship between average fragment size and sequence divergence. The level of sequence identity between insertion and
organellar DNA was calculated using the BESTFIT algorithm of the GCG package, which considers both nucleotide exchanges and
InDels (Devereux et al. 1984). Average fragment sizes were obtained from columns 2 and 4 of Table 1. Squares indicate long continuous integrants; open circles symbolize complex insertions derived from one organelle; and shaded
circles stand for complex insertions, including sequences from both organelles. (C) Intra-insertion patterns of sequence divergence. Sequence identity between NUPTs and ptDNA and between NUMTs and mtDNA was
assessed for the five largest complex loci as in B. NUPTs and NUMTs larger than 1 kb were considered. Insertion 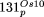 served as control. It consists of one large continuous NUPT subdivided into sets of fragments with sizes ranging from 20
kp to 1 kb; the level of its overall sequence identity is indicated by a gray line. In
served as control. It consists of one large continuous NUPT subdivided into sets of fragments with sizes ranging from 20
kp to 1 kb; the level of its overall sequence identity is indicated by a gray line. In 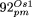 , NUPTs are indicated by continuous lines and NUMTs by dotted lines. Note that the more diverged fragments in
, NUPTs are indicated by continuous lines and NUMTs by dotted lines. Note that the more diverged fragments in 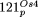 (26,331–31,363; 31,686–45,533; 46,267–53,135) are mostly derived from the IR region; in
(26,331–31,363; 31,686–45,533; 46,267–53,135) are mostly derived from the IR region; in 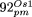 (55,367–56,927; 66,220–68,636; 75,288–81,711; 81,712–83,231) they originate from the IR region, and from positions 55,000–60,000
of ptDNA. The most diverged regions in
(55,367–56,927; 66,220–68,636; 75,288–81,711; 81,712–83,231) they originate from the IR region, and from positions 55,000–60,000
of ptDNA. The most diverged regions in 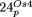 (10,721–16,939; 16,940–20,685) derive again from IR sequences.
(10,721–16,939; 16,940–20,685) derive again from IR sequences.











