
Structures of the complex integrants 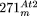 and
and 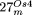 , which contain rearranged DNA from the mitochondrial chromosome of Arabidopsis and rice, respectively. (A) Structure of the Arabidopsis (At-mt) and rice (Os-mt) mtDNAs. For the Arabidopsis mtDNA, the position of the four single-copy regions A, B, C, and D, as well as the three pairs of specific repeats (I, II,
and III), are presented. Repeats I (positions 44,698–48,894 and 178,863–183,059) and II (103,805–104,337 and 227,087–227,619)
are directly oriented, while the two repeat III sequences (112,147–118,736 and 297,580–290,991) are inverted. The portion
of the mitochondrial chromosome included in each nuclear insertion (bottom) is indicated by black lines (as in the previous figures). (B) Structure of
, which contain rearranged DNA from the mitochondrial chromosome of Arabidopsis and rice, respectively. (A) Structure of the Arabidopsis (At-mt) and rice (Os-mt) mtDNAs. For the Arabidopsis mtDNA, the position of the four single-copy regions A, B, C, and D, as well as the three pairs of specific repeats (I, II,
and III), are presented. Repeats I (positions 44,698–48,894 and 178,863–183,059) and II (103,805–104,337 and 227,087–227,619)
are directly oriented, while the two repeat III sequences (112,147–118,736 and 297,580–290,991) are inverted. The portion
of the mitochondrial chromosome included in each nuclear insertion (bottom) is indicated by black lines (as in the previous figures). (B) Structure of 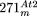 and
and 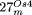 , depicted according to the scheme used in Figures 1 and 2. For
, depicted according to the scheme used in Figures 1 and 2. For 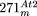 , small NUMTs are indicated by Arabic numerals and the positions of their homologs in the Arabidopsis mtDNA are as follows: 1 (170,597–170,530); 2 (170,530–170,597); 3 (191,246–191,300); 4 (25,397–25,647); 5 (28,644–28,687);
6 (279,338–279,090); 7 (280,258–279,809); and 8 (191,552–192,070). Four major rearrangements can be recognized (see text).
(1) A 5.5-kb region, derived from the D-region, is duplicated (positions 1–5545 and 47,106–52,650) in
, small NUMTs are indicated by Arabic numerals and the positions of their homologs in the Arabidopsis mtDNA are as follows: 1 (170,597–170,530); 2 (170,530–170,597); 3 (191,246–191,300); 4 (25,397–25,647); 5 (28,644–28,687);
6 (279,338–279,090); 7 (280,258–279,809); and 8 (191,552–192,070). Four major rearrangements can be recognized (see text).
(1) A 5.5-kb region, derived from the D-region, is duplicated (positions 1–5545 and 47,106–52,650) in 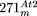 . The two sequences are more similar to each other than to mtDNA, and both harbor a characteristic 68-bp insertion derived
from region C (designated as “1” and “2”), suggesting that the insertion of this specific mtDNA segment in the nucleus occurred
before the duplication. (2) A 2.6-kb region (positions 5546–8207), derived from region C, is inserted between the D′ terminus
of the mtDNA insertion (see above) and the 5.5-kb duplication described above. (3) A 1.8-kb stretch, containing a structure
of six short NUMTs, together with very short stretches of nonorganelle DNA, is present at positions 74,295–76,084. The six
NUMTs derive from all four major regions of the mtDNA (including those absent at the A/D junction: see text and below). The
1.8-kb insertion is flanked by a duplication of 9 bp (CTTTACGAG) present in the D region, implying that it was inserted after
formation of the D′–A′–C–B structure. (4) At positions 129,022–129,597, between A region and III-type repeat, a short stretch
of the B region is found. This could be the result of imprecise homologous recombination between repeat III sequences, affecting
the B region adjacent to one of the repeats. Alternatively, this short region might represent the beginning of the 350-kb
duplication that was previously detected in
. The two sequences are more similar to each other than to mtDNA, and both harbor a characteristic 68-bp insertion derived
from region C (designated as “1” and “2”), suggesting that the insertion of this specific mtDNA segment in the nucleus occurred
before the duplication. (2) A 2.6-kb region (positions 5546–8207), derived from region C, is inserted between the D′ terminus
of the mtDNA insertion (see above) and the 5.5-kb duplication described above. (3) A 1.8-kb stretch, containing a structure
of six short NUMTs, together with very short stretches of nonorganelle DNA, is present at positions 74,295–76,084. The six
NUMTs derive from all four major regions of the mtDNA (including those absent at the A/D junction: see text and below). The
1.8-kb insertion is flanked by a duplication of 9 bp (CTTTACGAG) present in the D region, implying that it was inserted after
formation of the D′–A′–C–B structure. (4) At positions 129,022–129,597, between A region and III-type repeat, a short stretch
of the B region is found. This could be the result of imprecise homologous recombination between repeat III sequences, affecting
the B region adjacent to one of the repeats. Alternatively, this short region might represent the beginning of the 350-kb
duplication that was previously detected in 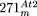 , but not sequenced (Stupar et al. 2001). For
, but not sequenced (Stupar et al. 2001). For 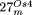 , the two large insertions of nonorganelle DNA into nuclear DNA of organellar origin are designated as “i1” and “i2” (Supplemental
Table 1). (C) Rearrangements of the Arabidopsis mtDNA due to recombination across repeat regions. The four single-copy regions A, B, C, and D are separated by two pairs
of repeats (I: direct repeats, III: inverted repeats); recombinations involving repeat II sequences are not shown because
they are not relevant for the generation of
, the two large insertions of nonorganelle DNA into nuclear DNA of organellar origin are designated as “i1” and “i2” (Supplemental
Table 1). (C) Rearrangements of the Arabidopsis mtDNA due to recombination across repeat regions. The four single-copy regions A, B, C, and D are separated by two pairs
of repeats (I: direct repeats, III: inverted repeats); recombinations involving repeat II sequences are not shown because
they are not relevant for the generation of 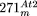 . Recombination across repeat II sequences results in A–D and B–C circles, while the A–B–D′–C′ structure derives from III–III
recombination (D′ and C′ refer to inverted D and C sequences, respectively). The A–D–B′–C′ arrangement originates from the
normal A–B–C–D structure by two rounds of recombination; the first, between the pair of repeat III sequences, inverts the
orientation of one of the repeat I sequences, allowing a second recombination between the now inverted repeat I sequences,
resulting in an A–D–B′–C′ circle. Another alternative structure, A–C′–B–D′, has been described before (Klein et al. 1994; Unseld et al. 1997). (D) Origin of
. Recombination across repeat II sequences results in A–D and B–C circles, while the A–B–D′–C′ structure derives from III–III
recombination (D′ and C′ refer to inverted D and C sequences, respectively). The A–D–B′–C′ arrangement originates from the
normal A–B–C–D structure by two rounds of recombination; the first, between the pair of repeat III sequences, inverts the
orientation of one of the repeat I sequences, allowing a second recombination between the now inverted repeat I sequences,
resulting in an A–D–B′–C′ circle. Another alternative structure, A–C′–B–D′, has been described before (Klein et al. 1994; Unseld et al. 1997). (D) Origin of 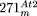 . The structure of
. The structure of 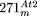 is depicted as a circle around mtDNA of the A–D–B′–C′-type. The
is depicted as a circle around mtDNA of the A–D–B′–C′-type. The 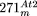 insertion contains the four major segments of mtDNA in the order D′–A′–C–B, whereby parts of D′ and A′ are either absent
in the nuclear mtDNA sequence, or form part of the 350-kb duplication for which no sequence information is available (Stupar et al. 2001). The deletion at the A/D junction is indicated by the dotted segment of the circle and contains mtDNA from position 183,060–209,
928 (region D), an entire I-type repeat, and the region extending from 343,608 to 44,697 (from region A) (see A). The breakpoint (indicated by an asterisk) that gave rise to the linear D′–A′–C–B structure should be located in region
D, close to the type III repeat; in fact, both ends of
insertion contains the four major segments of mtDNA in the order D′–A′–C–B, whereby parts of D′ and A′ are either absent
in the nuclear mtDNA sequence, or form part of the 350-kb duplication for which no sequence information is available (Stupar et al. 2001). The deletion at the A/D junction is indicated by the dotted segment of the circle and contains mtDNA from position 183,060–209,
928 (region D), an entire I-type repeat, and the region extending from 343,608 to 44,697 (from region A) (see A). The breakpoint (indicated by an asterisk) that gave rise to the linear D′–A′–C–B structure should be located in region
D, close to the type III repeat; in fact, both ends of 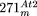 consist of sequences from the D region. Additional rearrangements of
consist of sequences from the D region. Additional rearrangements of 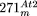 , in particular the duplication resulting in the 350-kb region identified before (Stupar et al. 2001), are indicated by arrows.
, in particular the duplication resulting in the 350-kb region identified before (Stupar et al. 2001), are indicated by arrows.











